A binary classifier makes decisions with confidence levels. Usually it’s imperfect: if you put a decision threshold anywhere, items will fall on the wrong side — errors. I made this a diagram a while ago for Turker voting; same principle applies for any binary classifier.

So there are a zillion ways to evaluate a binary classifier. Accuracy? Accuracy on different item types (sens, spec)? Accuracy on different classifier decisions (prec, npv)? And worse, over the years every field has given these metrics different names. Signal detection, bioinformatics, medicine, statistics, machine learning, and more I’m sure. But in R, there’s the excellent ROCR package to compute and visualize all the different metrics.
I wanted to have a small, easy-to-use function that calls ROCR and reports the basic information I’m interested in. For preds, a vector of predictions (as confidence scores), and labels, the true labels for the instances, it works like this:
> binary_eval(preds, labels)
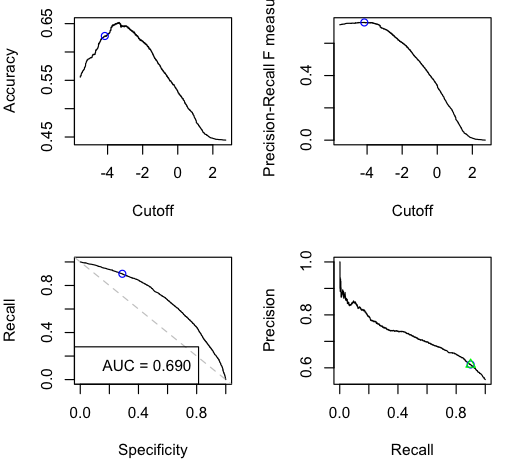
These are four graphs showing variation of classifier performance as the cutoff changes. There’s more than one because they convey different, interesting information. There are four because then they arrange nicely into a square. I wanted to balance having enough information vs. simplicity. The graphs are:
| Accuracy — how raw accuracy changes with different cutoffs. The extreme left and right sides are baselines of always guessing positive vs always guessing negative — so their accuracies are the respective population rates. (So you can see the data set is unbalanced; 55:45 pos:neg items.) | F-score — I want to kill this graph and replace it with something more useful. Kappa? But positionally it’s nice here: cutoff on the x-axis like the topleft, but it’s a precision/recall metric so goes along with the bottomright. |
| ROC aka Sensitivity-Specificity curve — Left side is low cutoff (aggressive), right side is high cutoff (conservative). Dashed line is for flipping weighted coins. “AUC” is area under curve which is what it sounds like. Is the Gini coefficient rescaled from [-1,+1] to [0,1]. Note that unlike all the other graphs, ROC & AUC are invariant across different item label proportions. (Given a trained classifier, precision and accuracy are influenced by the balanced-ness of the items; recall and specificity are not.) | Precision-Recall curve — Left side is high cutoff (conservative), right side is low cutoff (aggressive). Note this directionality is opposite of the 3 other panels. In particular, the recall axis is different than in the ROC plot. I debated flipping the axes, but prec vs rec is standard practice elsewhere. The green triangle is the point of best F-score. (Other possible points to highlight might be prec-rec break-even, or argmaxes for F2, or F0.5… all these prec-rec metrics seem like ad-hoc hacks to me.) |
The other thing binary_eval does is show one cutoff point across all the graphs — the blue circle — and display its associated performance metrics. So here, it used the naive cutoff (0 for real-valued predictions; 0.5 for probabilities) and plotted that point with the blue circle on all the graphs. Plus it reports for that cutoff:
Predictions seem to be real-valued scores, so using naive cutoff 0: Acc = 0.531 F = 0.338 Prec = 0.780 Rec = 0.216 Spec = 0.924 Balanced Acc = 0.570
So it’s very apparent that the naive cutoff isn’t performing well here. We can call it and supply a cutoff, or perhaps better, ask it to pick a cutoff that maximizes any metric that ROCR knows about. For example, calibration for accuracy or F-score:
| binary_eval(preds, labels, ‘acc’) | binary_eval(preds, labels, ‘f’) |

|
 |
I added the function to my R utilities file, here: dlanalysis/util.R.
Some people in machine learning like to use standalone programs like PERF to compute all these metrics. R is better if you know it. You can calculate all sorts of things with it. Above I was using R for the evaluation of the outputs of a command-line classifier, importing them easily with scan() and scan(pipe(“cut -f1 < data.svmlight_format”)).
The best Wikipedia page to start reading about the zillions of all of these interrelated confusion matrix metrics might be the the ROC page.
If you want to see lots and lots more evaluation metrics, look at the “extended classifier evaluation” section of LingPipe’s sentiment analysis tutorial. And once you get into multiclass and such, all complexity hell breaks loose.

This is an excellent overview of classifier evaluation. I wrote a posting, specifically on AUC, at:
ROC Curves and AUC
I’m loving R for this kind of stuff because of the graphics. It’s so much easier to understand that way than in a table of numbers.
I’d suggest (a) putting recall on the horizontal axis in both of the bottom graphs, (b) making the graphs square rather than rectangular, and (c) fixing axes to the range [0,1].
In a summary plot like this, I’d also like to see number of positive and negative examples in the gold standard, the name of the data set, name of classifier and any params that can be indicated. I always find myself fishing for that in papers. You might also indicate graphs are uninterpolated, though that’s obvious from looking at PR.
There’s no reason you couldn’t overlay multiple evals here in different colors, too.
I’d love to see posterior intervals in addition to single numbers in the plots. Wouldn’t a simple bootstrap work and be simple to implement in R? Of course, that’d get hopelessly messy and prone to misinterpretation with multiple evals on the same plot.
Given our focus on high recall, we’re often zooming in on the 95-100% recall section of the graph, but presumably that’s all settable as in the rest of R’s plotting functions.
Thanks for the LingPipe pointer — there’s also extensive discussion of classifier evaluation in our javadoc for ConfusionMatrix, PrecisionRecallEvaluation, and ScoredPrecisionRecallEvaluation.
Pingback: Text Classification for Sentiment Analysis – Precision and Recall «streamhacker.com
Hi Brendan,
a handy utility. However, have you left out a “last” function i.e.
> binary_eval(two.out.of.3.marker[C!="A"],Test2)
AUC = 0.807
Predictions seem to be real-valued scores, so using naive cutoff 0:
Error in binary_eval(two.out.of.3.marker[C != "A"], Test2) :
could not find function “last”
last<-function(x){ sort(x)[length(x)] }
seems to fix it and gives sensible plots so I am assuming that is all that last does
Bye
Rob
Rob: oops, sorry I forgot. Yup that’s it.